Problem
Toxicologic pathology is the study of the effects of compounds on biological beings, which includes pre-clinical studies that assess the safety and effectiveness of novel therapies during drug development, before clinical trials. Current regulations require that a novel therapeutic compound be administered to experimental animals that are subsequently sacrificed so that their tissues can be comprehensively examined for toxicologic effects, primarily in the liver and kidney. In each of these studies, tens to hundreds of experimental animals are used resulting in a very large number of tissue slides, most of which are completely normal. In the current workflow, all these slides must be manually through a highly complex visual examination to determine which slides are abnormal, An AI model to automate this would provide considerable resource savings in these studies, which would reduce the cost and time for novel drug development.
Approach
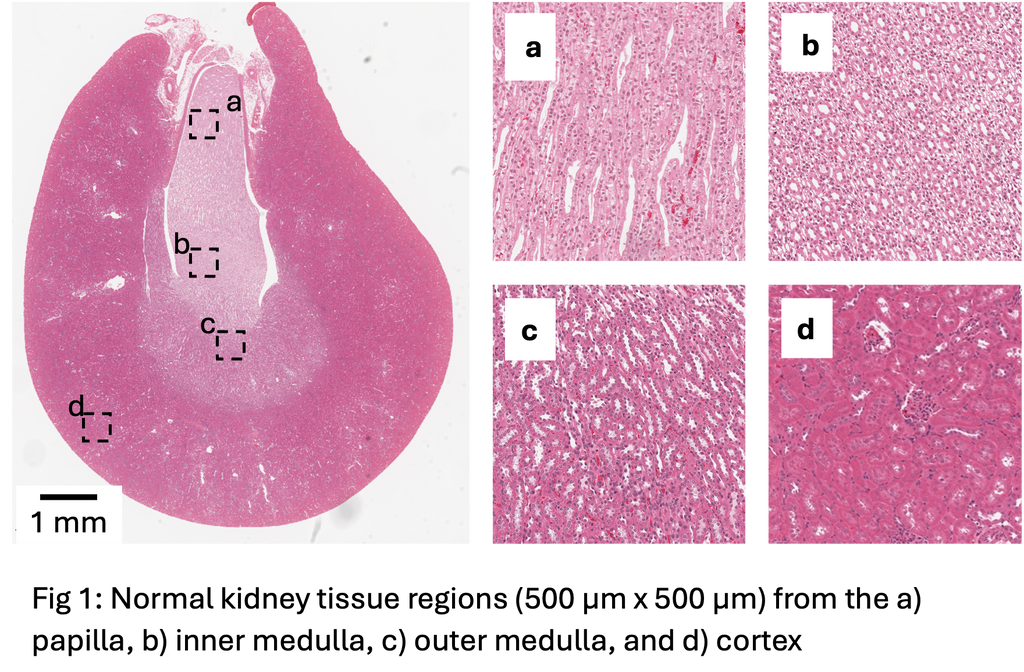
Previous work for kidney abnormality detection has shown that this is possible with an AUC of 0.74 when using a weakly supervised learning approach based on the UNI foundation model [1] for feature extraction and the CLAM approach [2] for classification. This is considerably lower than the performance we see in liver, which may be due to architectural complexity of kidney histology compared to the liver (Fig 1).
The first goal of this project is to evaluate whether newer and more powerful foundation models can achieve better performance through a benchmarking study on a subset of the data, which will establish a baseline for performance using off-the shelf methods.
The second goal is to create an enriched kidney structure map as a basis for classification, as this might improve tissue type recognition and ultimately abnormality detection. This would be done by applying a kidney histology segmentation model which was developed for human kidneys in another study within our group, then performing feature extraction and classification using that.
Data
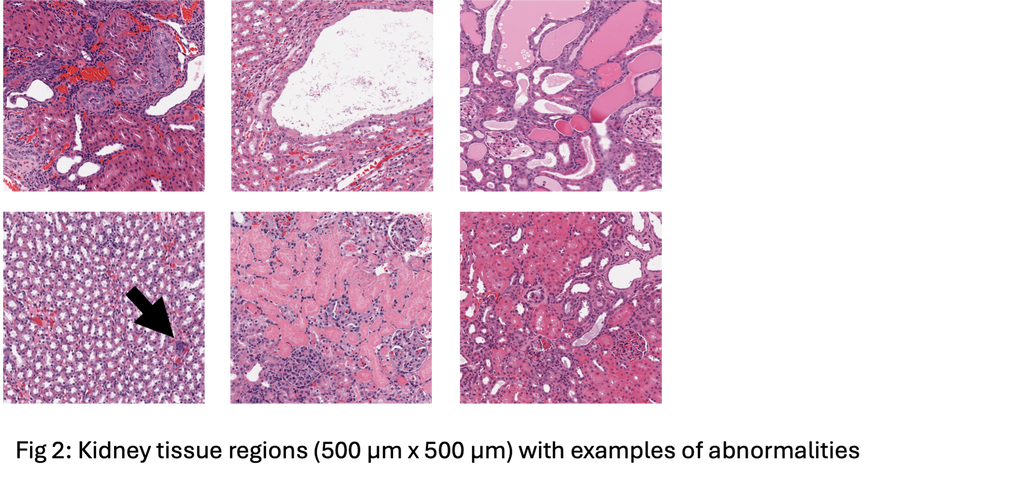
The data for this project comes from TG-GATES, a publicly available repository of compound safety studies. TG-GATES contains over 20,000 kidney digital pathology slides, organized by compound study, with labels indicating whether the slide has abnormalities (examples in Fig 2). We have a local clean copy of this data, so it will be immediately useable.
References
[1] Lu MY, Williamson DF, Chen TY, Chen RJ, Barbieri M, Mahmood F. Data-efficient and weakly supervised computational pathology on whole-slide images. Nature biomedical engineering. 2021 Jun;5(6):555-70. [2] Chen RJ, Ding T, Lu MY, Williamson DF, Jaume G, Song AH, Chen B, Zhang A, Shao D, Shaban M, Williams M. Towards a general-purpose foundation model for computational pathology. Nature medicine. 2024 Mar;30(3):850-62.
Requirements
- Students with a major in computer science, biomedical engineering, artificial intelligence, physics, or a related area in the final stage of master level studies are invited to apply.
- Affinity with programming in Python and familiar with the deep learning libraries.
- Interest in deep learning and medical image analysis.
Information
- Project duration: 6-9 months
- Location: Radboud University Medical Center, Nijmegen, the Netherlands.
- You will be part of the Diagnostic Image Analysis Group (DIAG), which is a 160-person team of researchers focused on researching and developing AI for medical imaging. Specifically, this position is within a subset of DIAG called the Computational Pathology Group (CPG), whose research focus is the analysis of histopathological slides. Both groups are highly supportive and welcoming, and frequently organize various social activities that you could take part in.
- Note that per our union rules, Radboudumc is unable to provide compensation for Master's thesis projects. Relatedly, we are also unfortunately unable to sponsor those that require a visa for the Netherlands for Master's thesis projects.
- For more information, please contact Salma Dammak.

