Problem
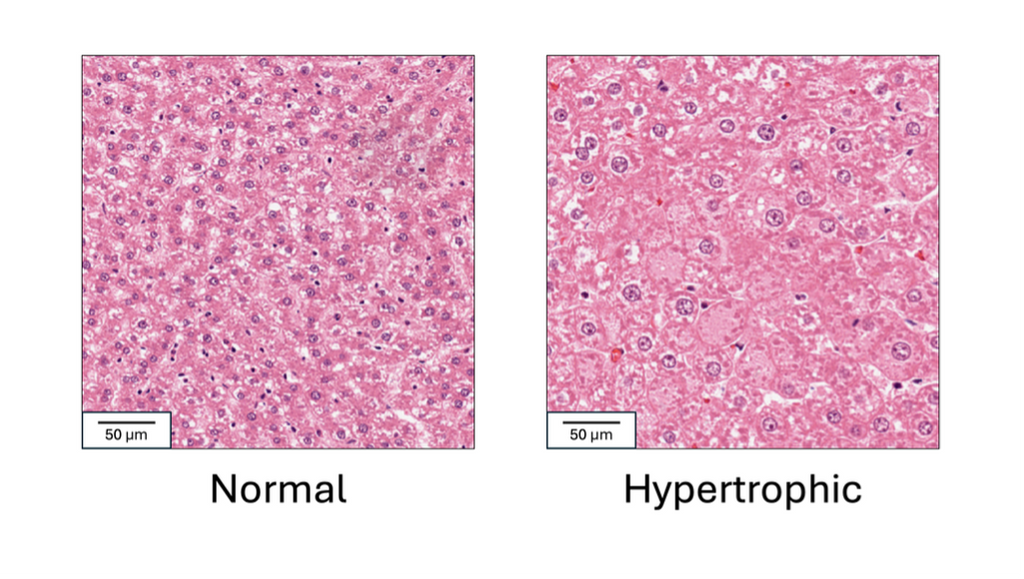
Drug safety studies are a key step in drug development and approval. In these studies, a group of experimental animals is given the target drug, and a control group is given a placebo or vehicle substance. Tissue samples from both groups are then microscopically analyzed by a pathologist to determine if any abnormalities are present in the target group. Hypertrophy, or cell enlargement, is a critical abnormality to detect in these studies, as it can indicate organ dysfunction, hormonal disruption or early carcinogenic effects. It often occurs in the liver, where normal cell size varies with age, strain, and diet, making it difficult to establish a universal threshold for hypertrophy. Therefore, to determine its presence, the pathologist compares tissues and various blood markers from the treated animals to those of the control animals. This is a challenging and highly time-consuming task, and the need for comparison can lead to missing hypertrophy when it is subtle. In this project, we aim to automatically detect liver hypertrophy on digital histology slides combined with blood markers to reduce the time and workload associated with drug development, which could allow drugs to reach those who need them sooner. We also aim to reduce the rate of missed hypertrophy, helping to prevent toxic drugs from advancing in the development pipeline.
Approach
Prior approaches for automatically detecting liver hypertrophy have analyzed individual slides in isolation, whereas pathologists rely on comparative analysis across multiple slides to detect hypertrophy. This single-slide approach may be prone to errors due to background variability, and in our internal testing showed a substantial drop in performance during external validation (from 0.83 AUC to 0.70 AUC). The first goal of this project is therefore to develop a new approach that can incorporate control group data when determining if a slide has hypertrophy, thereby aligning more closely with human pathology workflows and potentially improving detection accuracy.
In addition to comparing slides, pathologist use organ weight and blood markers to determine if hypertrophy is present. Prior work that incorporated this is very small in scale but shows promise. The second goal of this project to therefore to develop a method of incorporating multimodal data when determining if a slide has hypertrophy to further align with pathology workflows.
Data
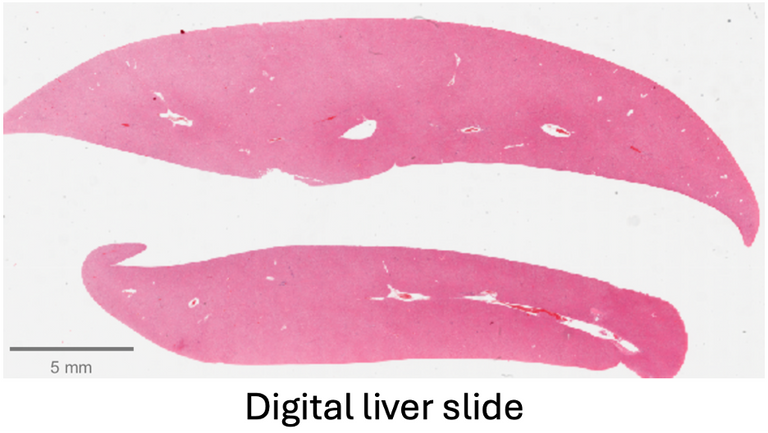
The data for this project comes from TG-GATES, a publicly available repository of rat drug studies. TG-GATES contains over 23,000 liver digital pathology slides, organized by drug study, with labels indicating whether each slide belongs to the control or treatment group and whether the slide has hypertrophy. Among these, more than 1,000 slides exhibit hypertrophy. Once a model is designed and validated with the TG-GATES, we also have access to a private 300-slide dataset that we can use for external validation. Both datasets also have organ weight information along with other markers.
Requirements
- Students with a major in computer science, biomedical engineering, artificial intelligence, physics, or a related area in the final stage of master level studies are invited to apply.
- Affinity with programming in Python, specifically in the PyTorch framework.
- Interest in deep learning and medical image analysis.
Information
- Project duration: 6-9 months
- Location: Radboud University Medical Center, Nijmegen, the Netherlands.
- You will be part of the Diagnostic Image Analysis Group (DIAG), which is a 160-person team of researchers focused on researching and developing AI for medical imaging. Specifically, this position is within a subset of DIAG called the Computational Pathology Group (CPG), whose research focus is the analysis of histopathological slides. Both groups are highly supportive and welcoming, and frequently organize various social activities that you could take part in.
- Note that per our union rules, Radboudumc is unable to provide compensation for Master's thesis projects. Relatedly, we are also unfortunately unable to sponsor those that require a visa for the Netherlands for Master's thesis projects.
- For more information, please contact Salma Dammak.

