Problem
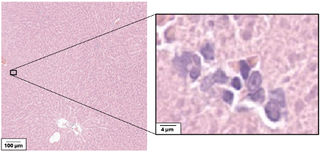
Drug safety studies are a key step in drug development and approval. In these studies, a group of experimental animals is given the target drug at different doses or for variable exposure times, in addition to a control group, which is given a placebo or vehicle substance. Tissue samples from all groups are then microscopically analyzed by a pathologist to determine if any abnormalities are present in the treated groups. One of the key abnormalities they assess is the presence of inflammation. Mononuclear immune cells (MIC) are immune cells that include lymphocytes and macrophages, and which can indicate chronic inflammation. This signals mild drug toxicity and may also predict the occurrence of more severe toxic effects at higher doses or longer exposure times. However, these cells are very small and tend to be sparsely distributed, making them easy to miss as shown in the main figure.
An automated model could improve reproducibility and sensitivity of preclinical safety readouts, reducing false negatives that could delay drug development. By doing so, such models could shorten the time required to identify toxic compounds, reduce the number of animals needed in studies, and lower the risk of unsafe drugs moving forward in the pipeline. This would directly accelerate drug development, improve patient safety, and contribute to more ethical and cost-effective preclinical research.
Related work
Inflammatory cell detection is an active research area in clinical pathology, particularly with AI models developed for the TIGER challenge. In contrast, inflammation is under-explored in preclinical research. As summarized in the table below, a few studies have addressed automatic detection of inflammation and immune infiltration in preclinical pathology, but most stop at coarse-level analysis such as slide- or patch-level classification. The goal of this project is therefore to adapt clinical pathology approaches to create a pixel-level MIC segmentation algorithm for preclinical pathology.
| Paper | Num WSIs | Animal | Organ | Target classes | Performance |
|---|---|---|---|---|---|
| Cheng et al. [1] | 2,194 (984 positive, TG-GATES) | Rat | Liver, kidney | Cellular infiltration presence (binary) | Patch AUC > 0.98 |
| Yan et al. [2] | NR (20,709 tiles) | NR | Liver | 5 classes in NAFLD incl. inflammation | F1: 0.87 |
| Kuklyte et al. [3] | 1,342 in study, NR subset for this task | Rat | Liver, kidney, lung, heart | Inflammation, Immune cell infiltration | F1 score per organ: 0.45-0.95 |
| Baek et al. [4] | 201 in study, NR subset for this task | Rat | Liver | Inflammation, Immune cell infiltration | mAP: 96% for inflammation and 94% for infiltration |
NR = Not Reported
Approach
The research question is: Can AI-based segmentation models reliably detect and segment MIC clusters in rat liver slides from drug safety studies?
Which will be addressed through the following objectives:
- Benchmark existing nuclei segmentation and detection approaches on preclinical slides
- Develop or adapt a model for MIC cluster segmentation
- Validate performance against expert annotations using appropriate segmentation metrics (e.g., Dice, precision, recall)
Examples of off-the-shelf approaches to start with include Pathology-nnUnet, which was developed by our group, and HoVerNet, which is available through the publicly available TIAToolbox.
Data
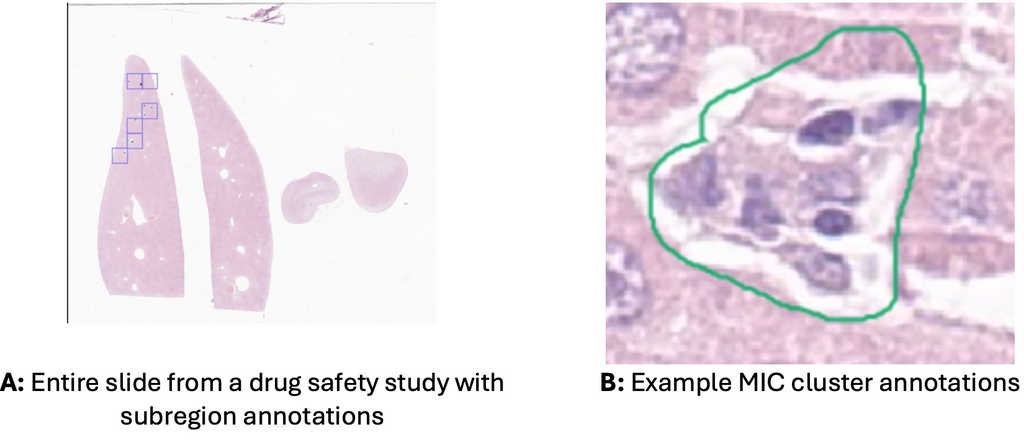
We currently have segmentations of MIC clusters in 86 H&E-stained pathology slides and expect another 43 within a few weeks. Obtaining exhaustive manual contours for this problem is highly time-consuming, so we opted to get contours within subregions of the slides as shown in A. For each slide, the pathologists selected six 1 mm x 1 mm subregions, and in each searched exhaustively for MICs. As MICs tend to group in clusters, they created an annotation that contained the entire cluster as shown in B. Additionally, if a look-alike cell or cell cluster was identified, it was also annotated as a counter example.
References
[1] Cheng, Z., Dai, W., & Sun, J. (2024, October). Visual Language Model for Preclinical Toxicologic Liver Histopathology Assessment. In Proceedings of the First International Workshop on Vision-Language Models for Biomedical Applications (pp. 41-48).
[2] Yan, R., He, Q., Liu, Y., Gou, J., Sun, Q., Zhou, G., He, Y. & Guan, T. (2022, October). DEST: deep enhanced swin transformer toward better scoring for NAFLD. In Chinese Conference on Pattern Recognition and Computer Vision (PRCV) (pp. 204-214). Cham: Springer Nature Switzerland.
[3] Kuklyte, J., Fitzgerald, J., Nelissen, S., Wei, H., Whelan, A., Power, A., Ahmad, A., Miarka, M., Gregson, M., Maxwell, M. Raji, R., Lenihan, J., Finn-Moloney, E., Rafferty, M., Cary, M., Barale-Thomas, E., & O’Shea, D. (2021). Evaluation of the use of single-and multi-magnification convolutional neural networks for the determination and quantitation of lesions in nonclinical pathology studies. Toxicologic Pathology, 49(4), 815-842.
[4] Baek, E. B., Hwang, J. H., Park, H., Lee, B. S., Son, H. Y., Kim, Y.B., Jun, S.Y., Her, J., Lee, J. & Cho, J.W. (2022). Artificial intelligence-assisted image analysis of acetaminophen-induced acute hepatic injury in Sprague-Dawley rats. Diagnostics, 12(6), 1478.
Requirements
- Master’s student in computer science, biomedical engineering, artificial intelligence, physics, or a related field
- Affinity with programming in Python, specifically in the PyTorch framework.
- Interest in deep learning and medical image analysis.
Information
- Project duration: 6-9 months
- Location: Radboud University Medical Center
- You will be part of the Diagnostic Image Analysis Group (DIAG) and Computational Pathology Group, whose research focus is the analysis of histopathological slides with deep learning techniques.
- You will have access to and work with a large GPU cluster.
- For more information, please contact Salma Dammak.

