Start date: 15-08-2020
End date: 15-02-2021
Clinical problem
Whole slide imaging technology allowed the digitization of conventional glass slides, which led to several new opportunities in the Pathology field, such as the integration of computational systems, most notably artificial intelligence (AI). However, the digitization process also brought along new challenges for the automated analysis of digitized images. One of the most important challenges arises from the presence of image artifacts. The image below shows examples of common types of artifacts found in whole slide images; (a) out-of-focus, (b) tissue folds, (c) ink, (d) dust, (e) pen mark, and (f) air bubbles.
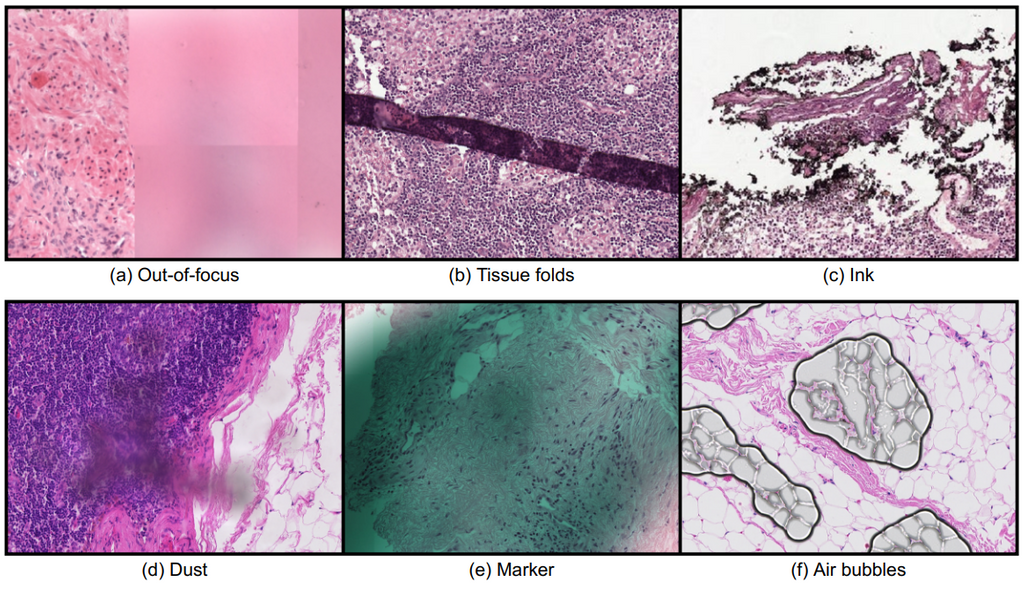
Depending on the severity of artifacts that are present, tissue regions that are important for diagnosis could be unclear, unusable, or even be completely missing. As a result, an AI system can fail to make a correct diagnosis in those regions. A quality control mechanism is needed to ensure that whole slide images are of good enough quality to be further analyzed by pathologists. Additionally, the quality system could provide recommendations, such as rescanning the whole slide image or leaving out defect regions of the slide when making diagnoses. Here is an example image with artifacts causing an AI algorithm to make incorrect predictions;
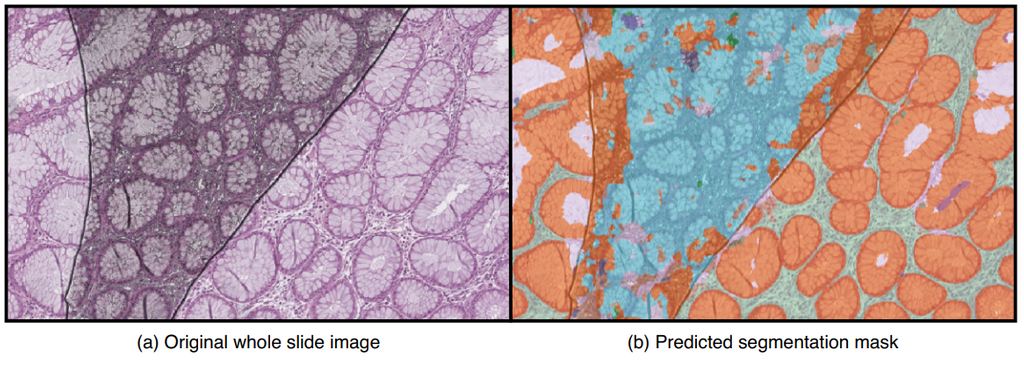
In the Pathology department of Radboudumc, every scanned whole slide image is manually inspected on its quality by technicians. Approximately 1.7% of 950 scanned slides are rescanned daily due to having too poor quality. The high volume of the slides and the sheer size of the scanned images (reaching up to 100,000 x 200,000 pixels) make manual control and supervision a highly time-consuming task for technicians.
Methods
We propose a solution to this problem in the form of an AI system that detects and highlights regions containing artifacts in whole slide images using deep learning methods. Additionally, the system provides a quality score for technicians to speed up the inspection process. The two main goals of this AI system are to speed up the process of digitization of slides and to make other AI algorithms more robust against artifacts.
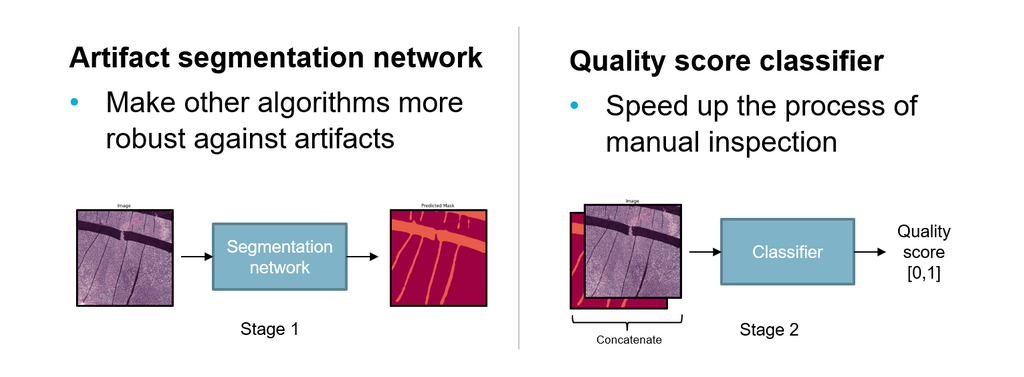
The system consists of two parts. First, an artifact segmentation network to make other AI algorithms more robust against artifacts. It does this by detecting and highlighting artifact regions in digitized images. AI algorithms can use the output segmentation mask to exclude artifact regions of an image during automated analysis. Second, a quality assessment network that helps to speed up the process of manual inspection by technicians. It does this by generating a quality assessment score for digitized images, ranging between 0 and 1, which indicates the overall quality. Using a threshold, technicians can choose how strict the quality assessment network filters out good and bad quality slides.
Results
We trained the artifact segmentation network on 142 slides, coming from various tissue types and stainings. The segmentation network achieved an overall Dice score of 0.870. Additionally, we evaluated the qualitative performance of the network on an external test set originating from a different medical center. The quality assessment network was trained 400 annotated slides with binary labels. This network achieved an overall AUC of 0.98.
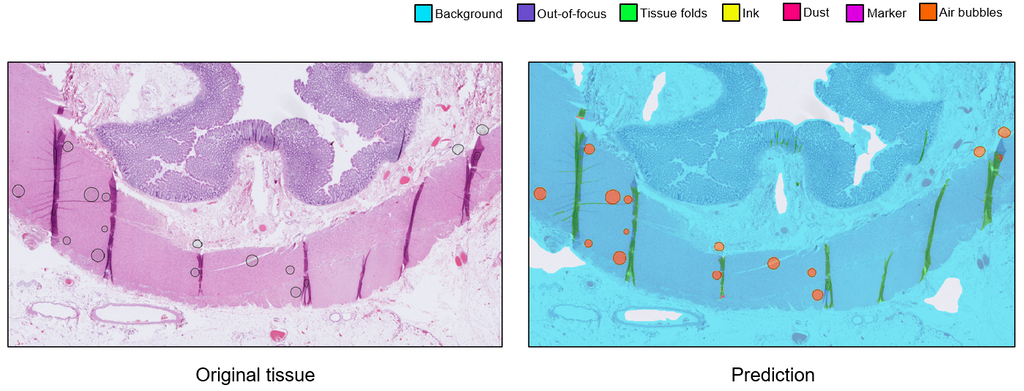
Conclusion
We developed an AI system consisting of an artifact segmentation network to make other algorithms more robust against artifacts and a quality assessment network to speed up the process of manual inspection. The generalization of both networks may be improved with more annotations. The average inference time per slide is ~6 minutes and can be done in parallel to reduce inference time. The system shows potential to be used in the AI-assisted future of the workflow in clinical practice. A possible next step of this project would be to implement the system in the daily workflow of pathologists and technicians.
Final report can be retrieved from here: MIDL short paper.
An interactive demo is accessible via Grand-Challenge:
The code for this project can be found in this GitHub repository.


